PhD students
|
|
Frullania genus is the most complex among liverworts, known for its rich chemical diversity. Its most distinctive compounds are sesquiterpene lactones and bibenzyls, but their biosynthetic pathways are mostly unknown. One of the first steps of terpenoids biosynthesis are catalyzed by terpene synthases. Recently, plant terpene synthases called microbial terpene synthase-like enzymes (MTPSLs) have been identified. My thesis will investigate terpene biosynthesis in Frullania liverworts, focusing on the functional and biochemical characterization of their MTPSLs. This includes assessing enzyme activity in heterologous systems like yeast and in planta, as well as in vitro, followed by detecting their products. I will also explore the structure-function relationship of these enzymes by introducing mutations in their conserved regions. Better understanding of liverwort terpenoid biosynthesis could offer insights into the evolutionary path of MTPSLs in non-seed plants. |
|
Nikolina Mrakovčić Jean Monnet University Saint-Étienne - France |
|
|
|
|
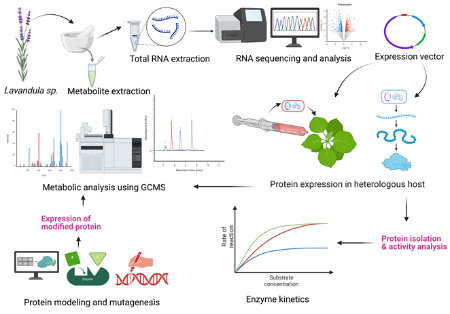
The genus Lavandula (lavender) consists of a diverse class of terpene synthases involved in plant defense, signaling, and therapeutic applications, as well as in the fragrance and cosmetics industries. My PhD objective is to characterize terpene synthases in Lavandula species based on transcriptomic data. |
Nirmal Yadav Jean Monnet University Saint-Étienne - France
|
|
The study will focus on determining the kinetic parameters of in vitro protein activity by expressing TPS in yeast and analyzing their product profiles through GC-MS. The project will integrate bioinformatics, molecular biology, and biochemical assays. | |
|
|
Terpenes are synthesized from the precursors isopentyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP), which are naturally derived from the methylerythritol phosphate (MEP) pathway or the mevalonate (MVA) pathway. This project aims to enhance terpenoid production through optimization of the MEP pathway with systems biology tools. The expression of the improved terpenoid pathway in combination with terpene synthases (TPS) will then create a highly efficient microbial production host. This host can then be adapted to different TPS and even previously engineered TPS to expand the terpene product range or enhance terpenoid production further. |
|
Nina Egeler Technical University of Munich Munich - Germany |
|
|
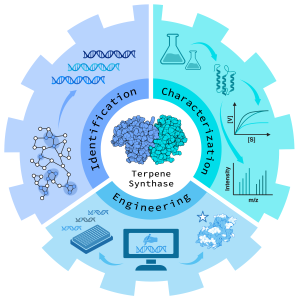
This project explores the potential of terpene synthases (TPS) for designing terpenoid scaffolds via a combination of computational and experimental methodologies. Specifically, it seeks to uncover new biocatalysts within the sesqui- and diterpene synthase families and investigate their physicochemical properties by integrating laboratory work and in silico analysis. |
|
|
This interdisciplinary approach focuses on: |
Grzergorz Jamróg Technical University of Munich Munich - Germany | |
|
|
Our group has deve-loped EnzyDock which tends to dock neutral and carboca-tionic ligands into en-zyme’s binding po-ckets. This tool would be used in ModBioTerp to un-derstand and im-prove TPSs: terpene synthases. |
|
|
Sumon Basak Bar Ilan University Ramat-Gan - Israel |
As a computational modeler, my responsibilities include integrating an ML-driven scoring function to enhance molecular docking accuracy and applying geodesic algorithms for transition state docking in Enzydock. I'll evaluate the efficient algorithms and develop parallel computing capabilities to accelerate simulations. | |
|
Additionally, I am responsible to develop a hybrid ML/MM potential and incorporate mutation capabilities using ML in EnzyDesign. The project would also involve designing enzyme variants such as CotB2 using PROSS, EnzyDock-EnzyDesign, and FuncLib. Through these efforts, my project would advance TPS engineering for applications in plant biotechnology. | ||
|
|
|
|
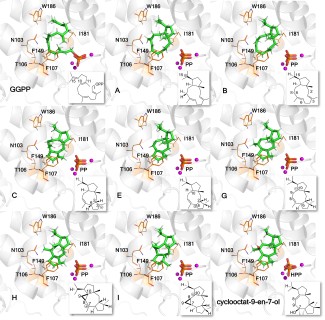
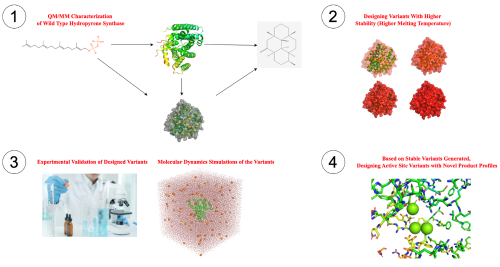
The aim of my project is to design diterpene synthases (diTPSs) with novel product profiles. My enzyme of interest is the Hydropyrene Synthase (Hp synthase). My objectives are: - Computational characterization of wild-type Hp synthase using QM/MM free energy simulations. |
Sangeet Dhiman Bar Ilan University Ramat-Gan - Israel
|
|
- Designing novel Hp synthase variants with enhanced stability and functionality using tools like PROSS. Methods: Multistate docking, multiscale modelling, molecular dynamics simulations, enhance sampling techniques, enzyme design tools, Python code development, EnzyDock code development. | |
|
|
The goal of my research is to model the mechanisms of terpene biosynthesis using deep learning methods. I will achieve this by assembling a dataset and designing deep learning models for predicting the reaction substrates and products directly from the amino acid sequences of terpene synthases. To this end, I will start with building a comprehensive database describing the reaction mechanisms of terpene synthases characterized to date. The database will contain all the required data for the deep learning models which will be designed followingly. I will then develop a deep learning model using state-of-the-art methods such as transformer neural networks for predicting the substrates, products, and reaction mechanisms of terpene synthases directly from their amino acid sequences. The model will consider vector representations of each protein generated by large protein language models, as well as structural domains detected in each protein structure. I expect this model to predict if a protein is a terpene synthase given its amino acid sequence. In the case of TPS, the model will also predict the details of the enzymatic activity, such as substrates, products, and reaction mechanisms. Last but not least, I will also be working on developing a generative algorithm for designing artificial TPSs with a desired function. |
|
Safa Mert Akmeşe Institute of Organic Chemistry and Biochemistry Prague - Czeck republic |
|
|
Terpenoid secondary metabo-lites are widely distributed in nature across diverse life forms and are generally utilized for defense and communication. They have many interesting industrial applications in phar-maceutics, cosmetics, in the food industry and others. |
|
|
In insects, TPSs are unrelated to their better-studied plant or microbial counterparts and evolved independently multiple times in several insect lineages. The focus of this project is the evolutionary origin of terpenoid biosynthesis in the western honeybee Apis mellifera, in which terpenoids are reported as pheromone components. We aim to uncover the terpenoid biosynthetic pathway in this species and to describe structural features of A. mellifera TPSs relevant for their enzymatic function. |
Monika Mikušková Institute of Organic Chemistry and Biochemistry Prague - Czech Republic | |
|
|
Terpene synthases (TPSs) produce a wide variety of natural compounds used in medicine, agriculture, and cosmetics, among other industries. However, predicting the products of a given TPS is challenging due to the diversity of their structures and mechanisms. Traditional methods for product prediction are time-consuming and often require a curated experimental approach for each novel enzyme.In contrast, state-of-the-art machine learning models offer a solution by leveraging structural and sequence features from previously characterized TPS enzymes. In my project, I combine data from structural conformations, sequences, substrate docking, and other computational simulations to identify subtle structural differences between TPS variants that result in specific products. These enhanced predictions of TPS specificity, based on structure-function relationships, can improve our understanding of terpene synthases and their potential for biotechnological applications. |
|
Szymon Szrajer University of Amsterdam Amsterdam - Netherlands |
|
|
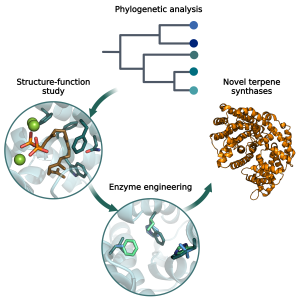
As part of the ModBioTerp project, I focus on evolutionary studies and the design of terpene synthases. These enzymes can synthesise a wide range of products, a trait linked to the complexity of their reaction mechanisms. The impact of protein structure on function remains largely unknown for terpene synthases, limiting their industrial applications. |
|
|
To address this issue, I use phylogenetic analysis to trace the evolution of enzymatic activity and assess the contribution of individual amino acids to product specificity. In addition to the biochemical characterisation of biocatalysts, I will perform in silico investigations involving docking experiments and molecular dynamics simulations. These insights will be used to guide engineering efforts and serve as a basis for the further development of enzyme variants. |
Victor Pricop KTH Royal Institute of Technology, Stockholm - Sweden | |












 Université Jean Monnet
Université Jean Monnet