Enzyme modelling courseTraining event 1 - Amsterdam - 3 to 15 March 2025
During this first event, all Partners and newly recruited DCs met, presented themselves and initiated direct personal contact for future collaboration. DCs elected a DC representative to the Supervisory Board.
The aim of the course held during this training was to train the DCs in the most recent enzyme modelling technologies using machine learning and advanced computational analysis. This entailed how to:
generate reliable three-dimensional structures of enzymes using advanced tools like Alphafold and homology modelling methods;
advanced docking tools for enzyme modelling like EnzyDock;
perform advanced reaction modelling using tools like QM/MM and enhanced sampling MD simulations;
phylogenetic analysis of enzymes;
apply advanced machine learning and cheminformatics tools to analyse enzyme simulations and predict enzyme structures and specificity;
apply generative AI and basic web-based tools like PROSS, FireProt, and FuncLib to design novel enzymes.
|
|
|
|
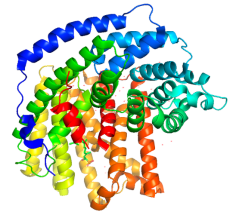
Model of 5-epi-aristolochene synthase from Nicotiana tabacum (Pymol, PDB database) |
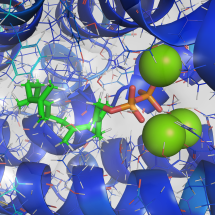
GGPP (substrate) Docking with Taxadiene Synthase (PDB: 3P5R) |
Students received in depth theoretical training during lectures and apply technologies during practical hands-on sessions. To deal with difference in previous exposure to computational techniques, pre-class reading material was used to provide a basic entry level, as well as advanced reading material for students with more advanced computational backgrounds.
In addition, depending on the topic, practical sessions were delivered in two modes, one low-code (mostly using online tools) as well as one focused on using command line/code). In addition to skills in modelling we also spend time on transferable skills to train DCs in collaboration between students with different academic backgrounds (in particular, computational background vs. experimental background).
|
|
|
|





 Université Jean Monnet
Université Jean Monnet